Results of allogeneic hematopoietic stem cell transplantation in a mixed cohort of patients with Ph-positive acute lympho-blastic leukemia
Accepted 17 February 2017
Published 10 March 2017
Summary
Additional chromosomal abnormalities (ACA) are rather common in Ph+ acute lymphoblastic leukemia (ALL). However, their prognostic significance in the era of protein tyrosine kinase inhibitors and allogeneic hematopoietic stem cell transplantation (allo-HSCT) is still poorly known. A recent study [1] has shown that ACA exert unfavorable effect upon HSCT results in adult patients with Ph+ALL.Patients and methods
We have performed a retrospective analysis of treatment results for a mixed cohort of the patients with Ph+ ALL, including 19 children (aged 5 – 18 y.o.) and 46 adults (aged 19 – 57 y.o.) who received allo-HSCT at our Institute over 2008 to 2015. Among sixty-five subjects with Ph+ ALL, the results of standard cytogenetic studies were available for 53 patients.Results
Thirty-three patients of 53 (51%) exhibited an isolated t(9;22) translocation. ACA were revealed in 20/53 patients (31%), including 13/53 (20%) subjects with 3 and more chromosome abnormalities. Chromosomes 1, 5, 7, 8, 9, 22 were most commonly affected with additional anomalies. Structural abnormalities attributable to ACA were imbalanced in 16 patients (80%), whereas only 4 patients (20%) showed balanced translocations. In a univariate analysis, significance was shown for the donor type (matched related and unrelated vs haploidentical, p=0.02), clinical stage at HSCT (1st remission vs other stages, p=0.01, for EFS only), additional chromosomal abnormalities (ACA-negative vs ACA-positive, p=0.04, for OS only), and, in particular, complex chromosomal aberrations (<3 anomalies vs ≥3 anomalies, p=0.01, for OS only). According to multivariate analysis, the number of additional chromosomal abnormalities per karyotype (HR 2.79, 95% CI 1.23-6.34; р=0.01, for OS only) and clinical stage at HSCT (HR 2.15, 95% CI 1.13-4.09; р=0.01, for EFS only) are independent prognostic factors for clinical outcomes.Conclusion
The study has shown that complex chromosomal anomalies and the stage of disease at the moment of HSCT are independent prognostic factors in a mixed cohort of Ph+ ALL patients treated with hematopoietic stem cell transplantation.Keywords
Acute lymphoblastic leukemia, Ph1-positive, allo-HSCT, additional chromosomal abnormalities.Introduction
Ph1-positive acute lymphoblastic leukemia (Ph+ALL) ranks a special place among lymphoid tumors. Initially, Ph1 positivity in ALL cells seemed an unexpected finding, since it challenged a specificity of t(9;22) translocation for chronic myeloid leukemia [17]. Afterwards, upon data gaining, it has become clear that Ph+ ALL may occur in all age groups, being more common in aged people, with minimal rates shown for children. Its prevalence is as high as 20% among adult ALL patients [15]. Until more recent times, the Ph+ ALL was considered an extremely unfavorable clinical variant with respect to all known treatment modes including hematopoietic stem cell transplantation (HSCT) [15]. This problem was solved as soon as different tyrosine kinase inhibitor drugs (TKIs) were introduced to clinics, thus enabling stable molecular remissions of the disease [3,6,13,18], and their usage for successful HSCT [2,16,23] including autologous (auto-) HSCTs [4,7,8,25]. ACA in Ph+ ALL are commonly seen, being described for 30-60% of the cases [5, 11, 14, 24]. Evaluation of allo-HSCT outcomes has shown that their prognosis depends on presence or absence of additional chromosome aberrations in cases of Ph1 chromosome positivity [1]. Moreover, it should be kept in mind that the ACA in Ph+ ALL patients could not be definitive. In particular, efficiency of chemotherapy and HSCT may be very good in cases of Ph1 chromosome combined with high hyperdiploidy [22]. We have previously confirmed it in our studies [9]. Such clinical aspect seems to be of high importance, in view of revisiting auto-HSCT for Ph+ ALL treatment [7,8,25].
Our study concerned a retrospective analysis of allo-HSCT performed in a mixed cohort of children and adults with Ph+ ALL who exhibited different clinical, transplant and cytogenetic characteristics.
Patients and Methods
The study was performed in a group of sixty-five patients with Ph+ ALL who underwent allo-HSCT at the First St. Petersburg I. Pavlov State Medical University over 2008 to 2015. We have used short-term 24-hour culturing of bone marrow cells without mitogen stimulation. Cytogenetic studies were carried out with a GTG chromosome staining using a standard technique [9]. Fluorescent in situ hybridization (FISH) with specific DNA probes proceeded according to the manufacturers’ protocols (MetaSystems, Germany; CytoCell, Great Britain). Interpretation of chromosomal aberrations was performed in accordance with an International System for Human Cytogenetic Nomenclature [20]. Evaluation of overall survival (OS) and event-free survival (EFS) was carried out in the patients with different demographic and clinical characteristics including gender and age, donor type, clinical state at the HSCT, conditioning regimen, source of stem cells and number of stem cells transplanted. The overall survival (OS) has been determined as the time period passed since HSCT to the patient’s death (for any reasons), or until the last examination date. Event-free survival (EFS) was evaluated as a time from HSCT to any adverse event (non-achievement of remission post-transplant, relapse, or death for any reasons), or till the last examination date. Statistical evaluation was performed with digital package ‘R’, version 3.1.1. (The R Foundation for Statistical Computing, Vienna Austria 2012). Survival curves have been plotted, according to Kaplan-Meier analysis. The survival plots were compared by means of a log-rank test; confidence levels by p<0,05 were considered significant. Multivariate analysis has been performed by the Cox regression model.
Results
The group of patients included 26 females (40%), and 39 males (60%), at the age between 5 and 48 years (a median of 26 years). Table 1 represents clinical and transplantation characteristics of the cohort under study. Thirty-one patients (48%) received allo-HSCT in the 1st remission, 20 subjects (31%) were transplanted in the 2nd remission, whereas 14 (21%) of the patients underwent allo-HSCT in active stage of the disorder. Bone marrow was a source of stem cells in 31 patients (48%), whereas peripheral blood stem cell transplants were used in 32 subjects (49%). Two patients got stem cells from the both sources. Reduced-intensity conditioning regimens (RIC) were used in 36 cases (55%), myeloablative treatment, in 29 cases (45%). Eighteen patients (28%) had HLA-identical sibling in their families; whereas 42 patients (65%) could be transplanted from HLA-matched unrelated donors found in an international registry. In 5 cases (7%), a related haploidentical HSCT was performed, since HLAmatched donor was absent in the family or blood donor Registries.
Table 1. Clinical and transplantation characteristics of Ph+ ALL patients under study
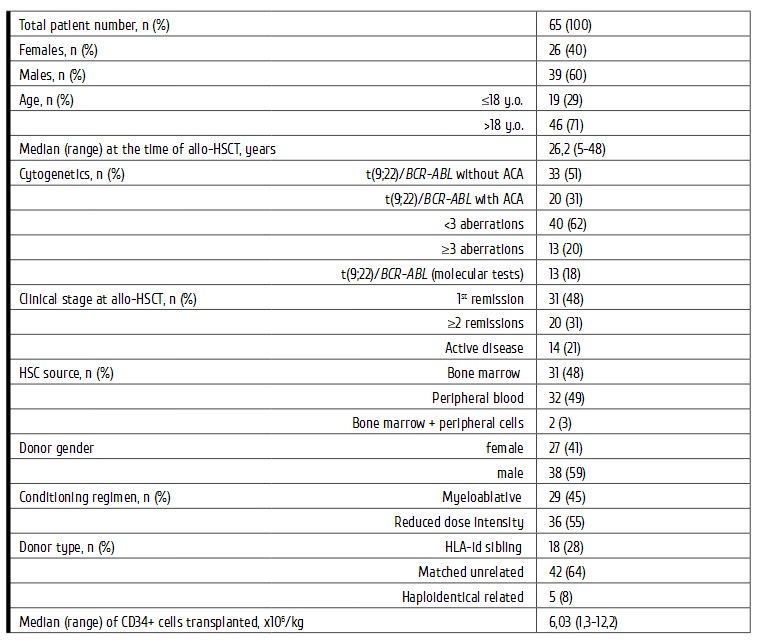
Cytogenetic characteristics of Ph-positive ALL
Primary diagnosis of Ph+ ALL was established by a standard cytogenetic examination performed in 53 (80%) of the patients. In 12 subjects (18%), the diagnosis was based on a fusion chimeric BCR-ABL gene found by FISH technique and chimeric bcr/abl transcript (р190 and/or p210) revealed with PCR.
Afterwards, we considered karyotypic changes in 53 patients with described pre-transplant cytogenetics. A t(9;22) (q34;q11) translocation, as sole karyotypic aberration, has been revealed in 33 patients, including 9 children (38%) and 24 (62%) adults. Accessory chromosomal anomalies were found in 20 patients, 5 (33%), in children and 15 (67%) in adults. The cytogenetic data for the 20 patients with additional chromosomal aberrations (ACA) are represented in Table 2. Both quantitative and structural ACA have been detected. Numerical abnormalities beared, mainly, on the chromosomes 1, 7, 8, 9, 10, being found in 12 out of 20 cases (60%). Trisomy 1 was nonrandom, being found in 2 patients (No5, 6). Similar repeated findings were made for trisomy10 (No4, 20), trisomy 22 (No4, 16), monosomy 7 (No8, 12). Trisomy 2 (in No20), trisomy 17 and 19 (No2), monosomy 9 (No16) were revealed in single patients.
Table 2. Karyotypes of the patients with Ph+ ALL with additional chromosomal abnormalities (ACA)
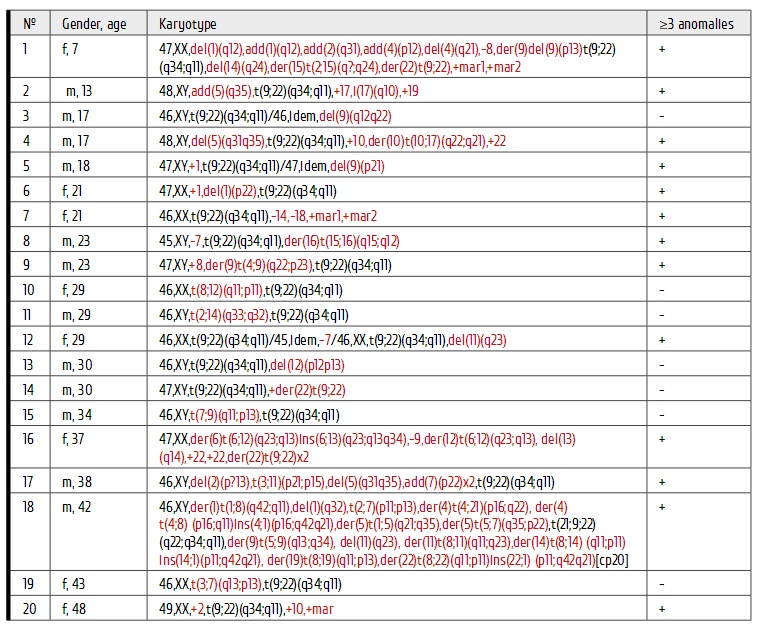
Note: additional chromosomal abnormalities are marked red.
Additional structural aberrations [except of t(9;22)] were registered in 20 patients. These were unbalanced in 16 of 20 cases (80%). Meanwhile, only 4 patients (20%) exhibited reciprocal translocations. Detailed evaluation of the chromosomal alterations, i.e., complete or partial monosomies and trisomies, is represented in Fig.1. Chromosomes 5, 7, 9, 2, 1, 17, 22 were most commonly involved into additional structural rearrangements. E.g., deletions and translocations in the short arm (p) of the chromosome 9 were observed in 4 patients out of 20 (No1, 5, 9, 15); reciprocal and unbalanced translocations with involvement of 7p were found in 3 cases (No17, 18, 19). Interstitial deletions and unbalanced translocations under involvement of 5q have been registered in 4 patients (No2, 4, 17, 18); deletions/translocations with chromosome 1, in 3 cases (No1, 6, 18); deletions and translocations of the chromosome 2, in 4 subjects (No1, 11, 17, 18). A structurally changed chromosome 17 was noted in 2 cases, as an isochromosome 17q (No2), or as a partner in unbalanced translocation (No4). An accessory derivate of the chromosome 22 was revealed in two patients (No14, 16). Unbalanced rearrangements involving other chromocomes occured occasionally.
A karyotype with three and more chromosomal aberrations was revealed in 13 patients (20%). As an example, the karyograms with a variant t(21;9;22)(q22;q34;q11) translocation and compound chromosomal abnormalities are presented on Fig. 2. The latter include ‘jumping’ segments (1q, 8q, 1q8q) to the partner chromosomes 1, 4, 5, 14, 19, 21 in a patient with chemoresistant Ph+ ALL (No18).
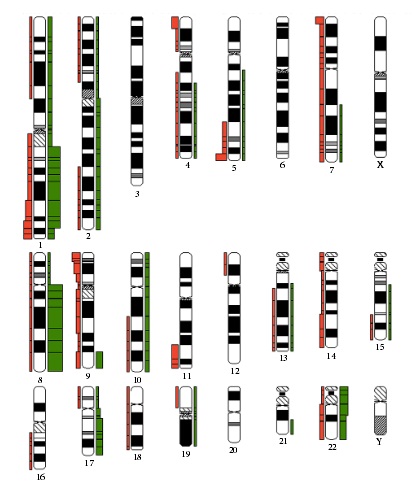
Figure 1. Figure illustrates the additional cytogenetic alterations in the cohort of Ph-positive ALL with the support of CYDAS (http://www.cydas.org/OnlineAnalysis/) [12]. Chromosomal gains are marked in green to the right, losses in red to the left. The thickness of the bars represents the number of cases showing the respective chromosomal gain or loss.
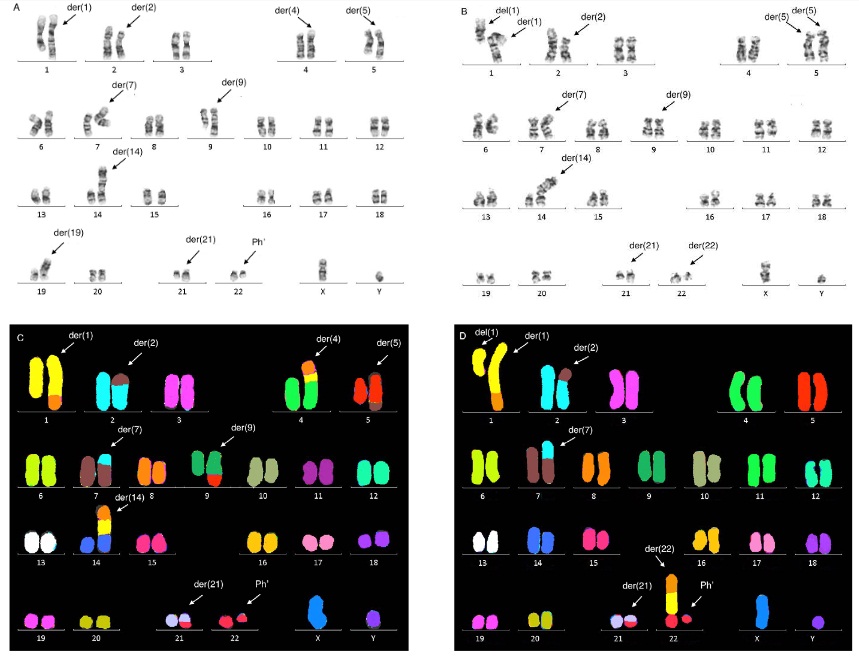
Figure 2. Karyograms of a bone marrow cells from a patient with chemoresistant Ph+ ALL and complex chromosomal abnormalities including a variant translocation t(21;9;22) and ‘jumping’ segments (1q, 8q, 1q-8q) to the partner chromosomes 1, 4, 5, 14, 21 (A,B – GTG banding; C,D, multi-coloured FISH).
Effects of cytogenetic, clinical and transplantation parameters upon clinical outcomes of allo-HSCT in the Ph+ALL patients
Furthermore, we performed an OS and EFS analysis in the patients different for their clinical and biological characteristics, e.g., gender, age, disease state, donor type, conditioning regimen, source of stem cells, presence of additional chromosomal aberrations and complex karyotypic abnormalities (≥3 per a karyotype). ///
A univariate analysis (Table 3) has shown significant differences in OS and EFS after allo-HSCT for the patients differing in their clinical stage at transplantation (for EFS only), donor type, as well as for groups with ACA, complex chromosomal aberrations (≥3 per karyotype), or ACA-free.
Table 3. Univariate analysis of overall survival (OS) and event-free survival (EFS) for the patients under study
Our data suggest that clinical efficiency was higher for transplantations performed at the 1st remission (for EFS only), in cases of matched related and unrelated donors, in absence of ACA in karyotype (for OS only), and, in particular, for complex chromosome aberrations (for OS only) (Fig. 3, 4). Meanwhile, no significant differences were obtained in the patients differing for their age, gender, conditioning regimens, stem cell source, time period from initial diagnosis to HSCT, and amount of transplanted CD34+ cells.
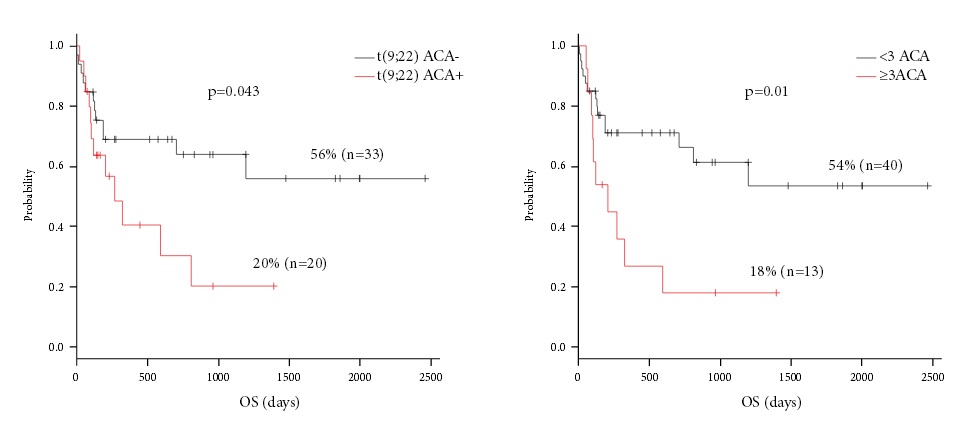
Figure 3. Overall survival after allo-HSCT in Ph+ ALL patients dependent on the presence of additional chromosomal abnormalities [except of t(9;22)] (left); and presence of ≥3 additional chromosomal aberrations (right).
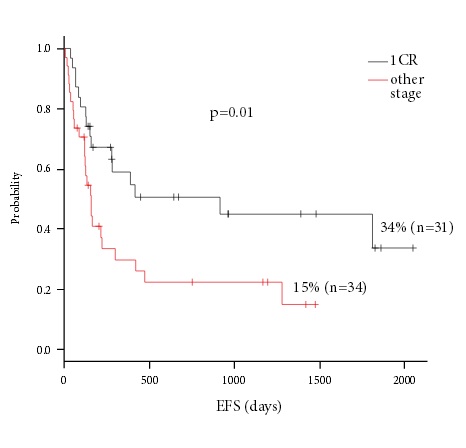
Figure 4. Event-free survival after allo-HSCT in Ph+ ALL patients dependent on clinical stage at time of HSCT.
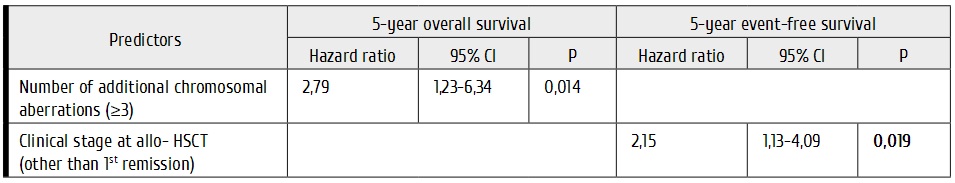
The results of multivariate analysis (Table 4) have shown that a presence of ≥3 chromosomal abnormalities in karyotype is an independent predictor for OS in ALL patients with t(9;22)/BCR-ABL translocation. Meanwhile, clinical stage at the time of allo-HSCT seems to be an independent predictor of event-free survival (EFS) of these patients.
Table 4. Multivariate analysis of survival predictors post-HSCT in Ph’-positive ALL
Discussion
Our study has confirmed recent conclusions on predictive significance of ACA in Ph+ALL patients [1]. According to several studies [5, 24], ACA in Ph+ ALL took place in 30-60% of these patients, and some of them underwent HSCT. As a rule, ACA affect chromosome pairs 7 to 9, 21 and 22 [11, 24]. Since advent of tyrosine kinase inhibitors (TKIs), predictive significance shifted towards ACA of chromosomes 9 and 22, making questionable a predictive role of other chromosomes. [21]. In our cohort, the structural changes in chromosomes 9 and 22 with 9p deletion and accessory Ph’-chromosome encountered in 6 patients. Moreover, ACA included also structural changes of chromosomes 5 (n=4), 7 (n=3), 1 (n=3), 17 (n=2) and some others. When discussing a role of ACA in Ph+ ALL patients, one should mention that some of them, by contrary, might lengthen OS and EFS. In particular, such situation is typical to the patients with a combination of one or two Ph1-cromosomes with high hyperdiploid karyotype [5, 19, 22]. In most of them, it was rather easy to achieve complete remission, due to positive predictive effect of high hyperdiploidy. Such patients were absent from our cohort. In the past, however, we have observed several ALL patients with two Ph’ chromosomes and high hyperdiploid karyotype who developed complete remission, even at standard chemotherapy, without TKI’s, with subsequent consolidation by means of autologous HSCT [9]. Some workers presume that only a combination of glucocorticoids and TKI’s is today sufficient to achieve a molecular remission, which could be fixed with auto-transplant [4, 7, 8]. As seen from our results, this cohort is heterogeneous, with respect to cytogenetic and prognostic parameters. Hence, appropriate therapy should be differentiated and based on cytogenetic data. In case of “favorable” combination of Ph’ chromosome and high hyperdiploid karyotype, the patients should be initially prepared by glucocorticoids and TKI’s (without application of ), then followed by auto-HSCT. What concerns more toxic protocols of therapy followed by auto-HSCT, they should be used, first of all, for treating the patients with other ACA types. Moreover, chemotherapy causes both de novo chromosome damage in Ph+ cells, and triggers a complex, prognostically adverse process of clonal evolution [10]. A resulting delay with HSCT in adult patients with Ph+ALL also remains unacceptable.
Conclusion
Our data show broad cytogenetic heterogeneity among leukemia patients with Ph+ ALL before HSCT. A significant subgroup exhibits additional chromosomal abnormalities which are associated with inferior clinical outcomes post-transplant. This phenomenon can be due to clonal evolution of malignant karyotype, or chemotherapy performed before HSCT, thus requiring further studies in the field.
Conflict of interest
All the authors have no conflict of interest to declare.
References
-
Aldoss I, Stiller T, Cao TM, Palmer JM, Thomas SH, Forman SJ, Pullarkat V. Impact of additional cytogenetic abnormalities in adults with Philadelphia chromosome-positive acute lymphoblastic leukemia undergoing allogeneic hematopoietic cell transplantation. Biol Blood Marrow Transplant 2015;21(7):1326-1329.
-
Cai WZ, Cen JN, Chen J, Fu CC, Han Y, Jin ZM, Ma X, Miao M, Qin HY, Tang XW, Xue SL, Sun AN, Chen SN, Wu DP. Major molecular response prior to allogeneic hematopoietic stem cell transplantation predicts better outcome in adult Philadelphia-positive acute lymphoblastic leukemia in first remission. Bone Marrow Transplant 2017;52(3):470-472.
-
Chiaretti S, Foa R. Management of adult Ph-positive acute lymphoblastic leukemia. Hematology 2015;2015:406-413.
-
Ding Z, Han MZ, Chen SL, MA QL, Wei JL, Pang AM, Zhang XY, Liang C, Yao JF, Cao YG, Feng SZ, Jiang EL. Outcomes of Adults with Acute Lymphoblastic Leukemia After Autologous Hematopoietic Stem Cell Transplantation and the Significance of Pretransplantation Minimal Residual Disease: Analysis from a Single Center of China. Chin Med J 2015;128(15):2065-2071.
-
Dombret H, Gabert J, Boiron JM, Rigal-Huguet F, Blaise D, Thomas X, Delannoy A, Buzyn A, Bilhou-Nabera C, Cayuela JM, Fenaux P, Bourhis JH, Fegueux N, Charrin C, Boucheux C, Lheritier V, Esperou H, Macintyre E, Vernant JP, Fiere D. Outcome of treatment in adults with Philadelphia chromosome-positive acute lymphoblastic leukemia – results of the prospective multicenter LALA-94 trial. Blood 2002;100(7):2357-2366.
-
Fielding AK, Rowe JM, Buck G, Foroni L, Gerrard G, Litzow MR, Lazarus H, Luger SM, Marks DI, McMillan AK, Moorman AV, Patel B, Paietta E, Tallman MS, Goldstone AH. UKALLXII/ECOG2993: addition of imatinib to a standard treatment regimen enhances long-term outcomes in Philadelphia positive acute lymphoblastic leukemia. Blood 2014;123(6):843-850.
-
Giebel S, Labopin M, Gorin NC, Caillot D, Leguay Th, Schaap N, Michallet M, Dombret H, Mohty M. Improving results of autologous stem cell transplantation for Philadelphia-positive acute lymphoblastic leukaemia in the era of tyrosine kinase inhibitors: A report from the Acute Leukaemia Working Party of the European Group for Blood and Marrow Transplantation. Eur J Cancer 2014;50(2):411-417.
-
Giebel S, Labopin M, Potter M, Poiré X, Sengeloev H, Socié G, Huynh A, Afanasyev BV, Schanz U, Ringden O, Kalhs P, Beelen DW, Campos AM., Masszi T, Mohty M, Nagler A. Comparable Results of Autologous and Allogeneic Hematopoietic Stem Cell Transplantation for Adult Patients with Philadelphia-Positive Acute Lymphoblastic Leukemia in First Complete Molecular Remission: An Analysis By the Acute Leukemia Working Party of the EBMT. Blood 2016;128(26):512.
-
Gindina TL, Mamaev NN, Barkhatov IM, Solomonova EV, Semyenova EV, Zubarovskaya LS, Morozova EV, Rudnitskaya YuV, Poppova MO, Alexeev SM, Uspenskaya OS, Bondarenko SN, Afanasyev BV. Complex chromosome changes in patients with recurrent acute leukemias after allogeneic hematopoietic stem cell transplantation. Ther Arkh 2012;84(8):61-66 [In Russia].
-
Gindina TL, Mamaev NN, Bondarenko SN, Semenova EV, Nikolaeva ES, Vlasova ME, Stancheva NV, Slesarchuk OA, Vavilov SN, Morozova EV, Alyanskiy AL, Afanasyev BV. Complex chromosomal aberrations in patients with post-transplantation replases of acute leukemias: clinical and theoretical aspects. Klin. Onkogematol. 2015;8(1):69-77.
-
Heerema NA, Harbott J, Galimberti S, Camitta BM, Gaynon PS, Janka-Schaub G, Kamps W, Basso G, Pui CH, Schrappe M, Auclerc MF, Carroll AJ, Conter V, Harrison CJ, Pullen J, Raimondi SC, Richards S, Riehm H, Sather HN, Shuster JJ, Silverman LB, Valsecchi MG, Arico M. Secondary cytogenetic aberrations in childhood Philadelphia chromosome positive acute lymphoblastic leukemia are nonrandom and may be associated with outcome. Leukemia 2004;18(4):693-702.
-
Hiller B, Bradtke J, Balz H, Rieder H. CyDAS: a cytogenetic data analysis system. Bioinformatics. 2005;21(7):1282-1283.
-
Kebriaei P, Saliba r, Rondon G, Chiattone A, Luthra R, Anderlini P, Andersson B, Shpall E, Popat U, Jones R, Worth L, Ravandi F, Thomas D, O’Brien S, Kantarjian H, de Lima M, Giralt S, Champlin R. Long-term follow-up of allogeneic hematopoietic stem cell transplantation for patients with Philadelphia chromosome-positive acute lymphoblastic leukemia: impact of tyrosine kinase inhibitors on treatment outcomes. Biol Blood Marrow Transplant 2012;18(4):584-592.
-
Ko BS, Tang JL, Lee FY, Liu MC, Tsai W, Chen YC, Wang CH, Sheng MC, Lin DT, Lin KH, Tien HF. Additional chromosomal abnormalities and variability of BCR breakpoints in Philadelphia chromosome/BCR-ABL-positive acute lymphoblastic leukemia in Taiwan. Am J Hematol 2002;71(4):291-299.
-
Moorman AV. The clinical relevance of chromosomal and genomic abnormalities in B-cell precursor acute lymphoblastic leukaemia. Blood Reviews 2012;26(3):123-135.
-
Parma M, Vigano C, Fumagalli M, Collnaghi F, Colombo A, Mottadelli F, Rossi V, Elli E, Terruzzi E, Belotti A, Cazzaniga G, Pogliani EM, Pioltelli P. Good outcome for very high risk adult B-cell acute lymphoblastic leukemia carrying genetic abnormalities t(4;11)(q21;q23) or t(9;22)(q34;q11), if promptly submitted to allogeneic transplantation, after obtaining a good molecular remission. Mediterr J Hematol Infect Dis 2015;7(1):e2015041.
-
Propp S, Lizzi FA. Philadelphia chromosome in acute lymphocytic leukemia. Blood 1970;36(3):353-360.
-
Ribera JM, Garcia O, Montesinos P, Brunet S, Abella E, Barrios M, Gonzales-Campos J, Bravo P, Hernandez-Rivas JM. Treatment of young patients with Philadelphia chromosome-positive acute lymphoblastic leukaemia using increased dose of imatinib and deintensified chemotherapy before allogeneic stem cell transplantation. Br J Haematol 2012;159(1):78-81.
-
Rieder H, Lufwig WD, Gassmann W, Maurer J, Janssen JW, Gokbudet N, Schwartz S, Thiel E, Loffler H, Bartram CR, Hoelzer D, Fonatsch C. Prognostic significance of additional chromosome abnormalities in adult patients with Philadelphia chromosome positive acute lymphoblastic leukaemia. Br J Haematol 1996;95(4):678-691.
-
Schaffer L., McGovan-Jordan J., Schmid M. ISCN. An international System for Human Cytogenetic Nomenclature. S. Karger, Basel. Switzerland, 2013, p.140
-
Short NY, Kantarjian HM, Sasaki K, Ravandi F, Ko H, Yin CC, Garcia-Manero G, Cortes JE, Garris R, O’Brien SM, Patel K, Khouri M, Thomas D, Jain N, Kadia TM, Daver N, Benton CB, Issa GC, Konopleva M, Jabbour E. Poor outcomes associated with +der(22)t(9;22) and -9/9p in patients with Philadelphia chromosome positive acute lymphobladstic leukemia receiving chemotherapy plus a tyrosine kinase inhibitor. Amer J Haematol. 2017;92(3):238-243.
-
Tauro S, McMullan D, Griffits M, Craddock C, Mahendra P. High-hyperploidy in Philadelphia positive adult acute lymphoblastic leukemia: case-series and review of literature. Bone Marrow Transplant 2003;31(9):763-766.
-
Wang L, Liu D-H. Tyrosine kinase inhibitor for treatment of adult allogeneic hematopoietic stem cell transplantation candidate with Philadelphia positive acute lymphoblastic leukemia. Chinese Medical Journal 2017;130(2):127-129.
-
Wetzler M, Dodge RK, Mrozek K, Stewart CC, Caroll AJ, Tantravahi R, Vardiman JW, Larson RA, Bloomfield CD. Additional cytogenetic abnormalities in adults with Philadelphia chromosome-positive acute lymphoblastic leukemia: a study of the Cancer and Leukemia Group B. Leukemia 2004;124(3):275-288.
-
Wetzler M, Watson D, Stock W, Koval G, Mulkey FA, Hoke EE, McCarty JM, Blum WG, Powell BL, Marcucci G, Bloomfield CD, Linker CA, Larson RA. Autologous transplantation for Philadelphia chromosome-positive acute lymphoblastic leukemia achieves outcomes similar to allogeneic transplantation: results of GALGB Sudy 10001 (Alliance). Haematologica 2014;99(1):111-115.
Accepted 17 February 2017
Published 10 March 2017


